Selects an optimal subset of channels to use as the common average reference
(CAR) for cortico-cortical evoked potential (CCEP) data, following the
CARLA (see 'Reference' and 'Citation'). Channels are ranked in
increasing order of their cross-trial covariance (or variance when only one
trial is available); subsets are then iteratively grown and the size that
yields the least anti-correlation between the candidate reference and the
remaining unreferenced channels is selected as optimal.
Usage
carla(
x,
nboot = 100L,
sensitive = FALSE,
min_size = NULL,
absolute_rank = FALSE,
virtual_reference = FALSE
)Arguments
- x
numeric array of shape
channels x time x trials; if a matrix is provided, it is treated aschannels x time(single trial). The signal must already be cropped to the responsive time window (typically0.01 sto0.3 spost-stimulus).- nboot
integer, number of bootstrapped trial resamplings used to estimate the optimization statistic; defaults to
100. Ignored when a single trial is supplied.- sensitive
logical; if
TRUE(and more than one trial is supplied), the more sensitive cutoff is used, corresponding to the channel count just before the first statistically significant decrease in the mean anti-correlation curve. The defaultFALSEreturns the global maximum of that curve.- min_size
integer, minimum subset size considered when
sensitive = TRUE; defaults tomax(2, ceiling(0.1 * nchan_good)), wherenchan_goodis the number of usable channels after the bad-channel mask has been applied (see Details). Floored at 2 because subset size 1 is never evaluated.- absolute_rank
logical; if
FALSE(the default), the per-channel ranking statistic is the signed mean of the off-diagonal trial-to-trial covariances, as in the original CARLA manuscript. This assumes a phase-locked evoked response, so trial-pair covariances on responsive channels are positive and a clean signed mean separates them from non-responsive channels. Set toTRUEto use the mean of the absolute covariances instead, which is more robust to evoked responses whose polarity flips across trials (e.g. alternating-polarity stimulation, biphasicCCEPswith jitter), at the cost of an upward bias on the non-responsive floor. Ignored when only one trial is supplied (the per-channel variance is used in that case).- virtual_reference
logical; if
TRUE, runs the modified CARLA before the iterative subset evaluation: rank channels once, designate the channel with the median rank as a "virtual reference", subtract its signal from every channel, then re-rank on the subtracted data. The virtual channel itself is pinned to the very end of the new ordering so it is never picked into the CAR until the final subset size. This is intended for data where the recording reference is contaminated by stimulation artifact or evoked activity: a contaminated reference makes every channel look artificially similar and biases the covariance/variance ranking statistic, but subtracting a mid-rank proxy of that contamination unbiases the ranking so genuinely responsive channels rise to the top. DefaultFALSEreproduces the original CARLA algorithm bit-for-bit.
Value
A list with the following elements:
channelsinteger vector, sorted indices (1-based) of the channels chosen to construct the common average reference.
carnumeric matrix of shape
time x trials(or numeric vector when only one trial is supplied) holding the common average signal computed fromchannels. Subtract this from each channel of the original signal to obtain the re-referenced data.orderinteger vector, indices of the good channels sorted in increasing order of the ranking statistic. Bad channels (zero variance / all-
NA; see Details) are excluded.varsnumeric vector of length
nchancontaining the per-channel ranking statistic (mean cross-trial covariance, or variance for a single trial). Bad channels keep their raw value (zero orNA) so callers can audit the mask.n_optimuminteger, the optimal subset size selected (indexes into
order).zmin_meannumeric matrix of shape
length(order) x nboot(or a length-length(order)vector for a single trial /nboot = 1) holding, for each subset size and bootstrap, the mean Fisher z-transformed correlation of the most globally anti-correlated unreferenced channel against the candidate CAR. Row 1 is alwaysNA(subset size 1 is not evaluated).bad_channelsinteger vector of channel indices that were excluded from the analysis because their ranking statistic was zero or
NA(flat / dead channels, plus the virtual channel itself whenvirtual_reference = TRUE).virtual_channelinteger, the index (1-based) of the channel used as the virtual reference when
virtual_reference = TRUE;NA_integer_otherwise.vars1numeric vector of the first-pass ranking statistic when
virtual_reference = TRUE(the post-subtraction statistic is returned invars);NULLotherwise.
Details
The function is a faithful port of the core CARLA.m routine; it does
not perform notch filtering, time-window cropping,
or grouping by stimulation site. Those steps belong to the surrounding
preprocess pipeline (see the example below).
For each candidate subset size \(n = 2, \ldots, N\), the
candidate reference is computed as the channel-wise mean of the \(n\)
lowest-ranked channels. Each of those \(n\) channels is then correlated,
in its unreferenced form, against every channel of the candidate
re-referenced subset. The resulting Pearson correlations are
Fisher z-transformed; the row corresponding to the most globally
anti-correlated channel is recorded as zmin. The optimal \(n\)
is the one that maximizes (i.e. makes least negative) the mean of
zmin.
Bad-channel mask. Channels whose ranking statistic is exactly
zero or NA are flagged as "bad" and excluded both from the CAR
candidate pool and from the target-channel correlation rows. This
catches flat / dead channels (constant signal, no usable variance) and
, when virtual_reference = TRUE, automatically removes the virtual
channel itself, since subtracting it from itself yields a zero trace.
All references to channel indices in the returned order,
n_optimum, and zmin_mean are with respect to the
good channels only; vars is full-length so callers can
inspect the raw scores.
References
The CARLA algorithm (virtual_reference = FALSE) is described in
doi:10.1016/j.jneumeth.2024.110153
; the modified CARLA precursor
(virtual_reference = TRUE) is described in
doi:10.1016/j.jneumeth.2025.110461
, with a reference implementation
at https://github.com/hharveygit/SPES_reference_contam. See
citation("ravetools") for the full bibliographic entries of
both manuscripts.
Examples
# ---- Simulate a small CCEP-like dataset --------------------------------
# 16 channels, 12 trials, sampled at 1 kHz, 0.5 s peri-stimulus epoch.
# Channels 1:4 are "responsive" (carry an evoked potential); the rest are
# noise-only and should make up the optimal CAR.
srate <- 1000
tt <- seq(-0.1, 0.4 - 1 / srate, by = 1 / srate) # time, seconds
nchan <- 16
ntrial <- 30
resp_ch <- 1:4
noise_ch <- 4:7
# Evoked potential template: damped sinusoid starting at t = 0
ep <- ifelse(tt >= 0,
80 * exp(-tt / 0.05) * sin(2 * pi * 12 * tt),
0)
# channels x time x trials
x_full <- array(rnorm(nchan * length(tt) * ntrial, sd = 5),
dim = c(nchan, length(tt), ntrial))
for (ch in resp_ch) {
for (k in seq_len(ntrial)) {
x_full[ch, , k] <- x_full[ch, , k] + ep * runif(1, -0.8, 1.2)
}
}
for (ch in noise_ch) {
for (k in seq_len(ntrial)) {
tmp <- x_full[ch, , k]
x_full[ch, , k] <- tmp + sign(ch %% 2 - 0.5) * 5 *
runif(length(tmp), 0.8, 1.2)
}
}
# Add artifacts common to all channels and trials
artifacts <- 6 * sin(2 * pi * 60 * tt) + 7 * sin(2 * pi * 24 * tt)
x_full <- sweep(x_full, 2L, artifacts, "+")
# ---- 1. Notch filter line noise (per channel, per trial) ---------------
# The CARLA paper notch-filters before ranking; the re-reference itself
# is applied to the original (unfiltered) signal.
x_clean <- x_full
for (ch in seq_len(nchan)) {
for (k in seq_len(ntrial)) {
x_clean[ch, , k] <- notch_filter(
x_full[ch, , k], sample_rate = srate,
lb = c(59, 119, 179), ub = c(61, 121, 181)
)
}
}
# ---- 2. Crop to the responsive window (0.01 s to 0.3 s post-stim) ------
resp_idx <- which(tt > 0.0 & tt <= 0.3)
x_resp <- x_clean[, resp_idx, , drop = FALSE]
# ---- 3. Run CARLA to pick reference channels ---------------------------
fit <- carla(x_resp, sensitive = TRUE, absolute_rank = TRUE,
virtual_reference = TRUE)
fit$channels # selected reference channels (should exclude 1:4)
#> [1] 5 6 7 8 9 10 11 13 14 15 16
fit$n_optimum # number of channels in the optimal CAR
#> [1] 11
# ---- 4. Re-reference the ORIGINAL (unfiltered) signal ------------------
# mean or median, your choice!
car_full <- apply(x_full[fit$channels, , , drop = FALSE], c(2, 3), mean)
# old-style: using all channels for CAR
car_old <- apply(x_full, c(2, 3), mean)
x_reref <- sweep(x_full, c(2, 3), car_full, "-")
x_compare <- sweep(x_full, c(2, 3), car_old, "-")
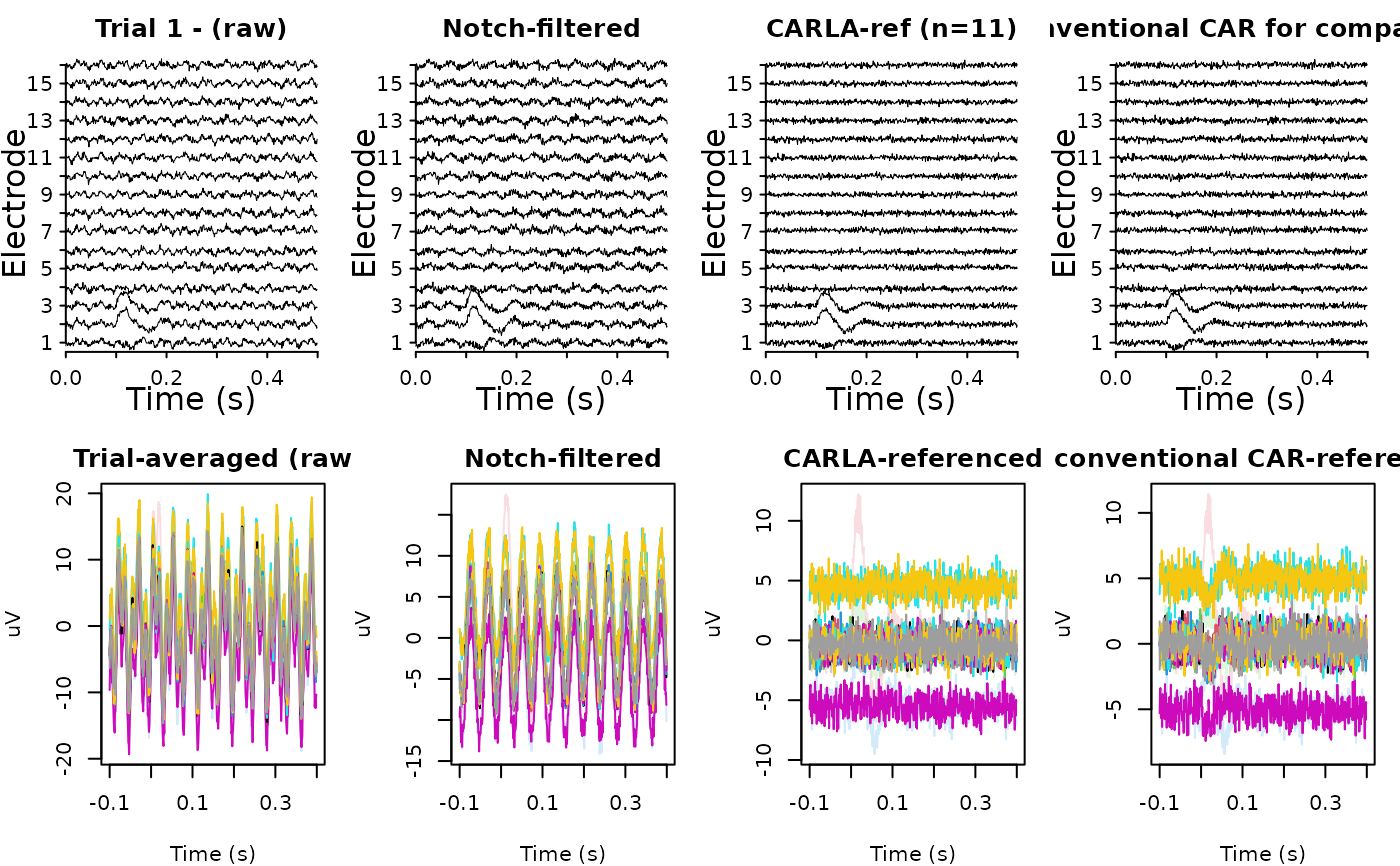
# ---- 5. Inspect: evoked potential is preserved on responsive channels --
op <- graphics::par(mfrow = c(2, 4), mar = c(4, 4, 2, 1))
ravetools::plot_signals(
signals = x_full[, , 1],
sample_rate = srate,
main = "Trial 1 - (raw)")
ravetools::plot_signals(
signals = x_clean[, , 1],
sample_rate = srate,
main = "Notch-filtered")
ravetools::plot_signals(
signals = x_reref[, , 1],
sample_rate = srate,
main = sprintf("CARLA-ref (n=%d)", length(fit$channels)))
ravetools::plot_signals(
signals = x_compare[, , 1],
sample_rate = srate,
main = "Conventional CAR for comparison")
col <- adjustcolor(seq_len(nchan))
col[resp_ch] <- adjustcolor(col[resp_ch], alpha.f = 0.2)
graphics::matplot(tt, t(rowMeans(x_full, dims = 2L)),
type = "l", lty = 1, xlab = "Time (s)", ylab = "uV",
main = "Trial-averaged (raw", col = col)
graphics::matplot(tt, t(rowMeans(x_clean, dims = 2L)),
type = "l", lty = 1, xlab = "Time (s)", ylab = "uV",
main = "Notch-filtered", col = col)
graphics::matplot(tt, t(rowMeans(x_reref, dims = 2L)),
type = "l", lty = 1, xlab = "Time (s)", ylab = "uV",
main = "CARLA-referenced", col = col)
graphics::matplot(tt, t(rowMeans(x_compare, dims = 2L)),
type = "l", lty = 1, xlab = "Time (s)", ylab = "uV",
main = "conventional CAR-referenced", col = col)
 graphics::par(op)
graphics::par(op)